Available and Ongoing Research Projects
The Kildea lab is able to offer research projects to strong undergraduate, MSc, and PhD students as well as postdocs. If interested, please contact John Kildea (john.kildea [at] mcgill.ca). Please note that due to the volume of inquiries received, it is not always possible to reply to all emails.
 |
|---|
| The Research Institute of the McGill University Health Centre |
Our research projects at the Research Institute of the McGill University Health Centre (RI-MUHC) fall into three broad programs:
- Neutron-Induced Carcinogenic Effects (NICE) projects,
- Radiation Oncology Knowledge Sharing (ROKS) projects, and
- Opal and the Quebec SmartCare Consortium projects.
Often, the ROKS and Opal projects are intertwined.
Students in the lab participate in the NSERC CREATE program Responsible Health and Healthcare Data Science that is run out of Université Laval in Quebec City. This offers access to various extra-curricular activities such as workshops and conferences that form nice line items on a CV.
The lab has a research associate (Luc) and research assistant (James) who help with all of the projects and we have access to the software development team of the Opal Health Informatics Group (John leads research and development for Opal) and resources in the Quebec SmartCare Consortium (a consortium of public institutions and private companies led by John). Also, we have access to national and international initiatives such as the Canadian Partnership for Quality Radiotherapy (until recently, John was vice chair) and the CodeX data standard accelerator at HL7 International (John represents the Canadian Organization of Medical Physicists on the operating committee). The senior students in the lab often mentor the junior students and help get projects running.
All projects in the Kildea lab are funded but all students are encouraged to and assisted in seeking scholarships as these make the funding go further and are also important CV line items.
Neutron-Induced Carcinogenic Effects
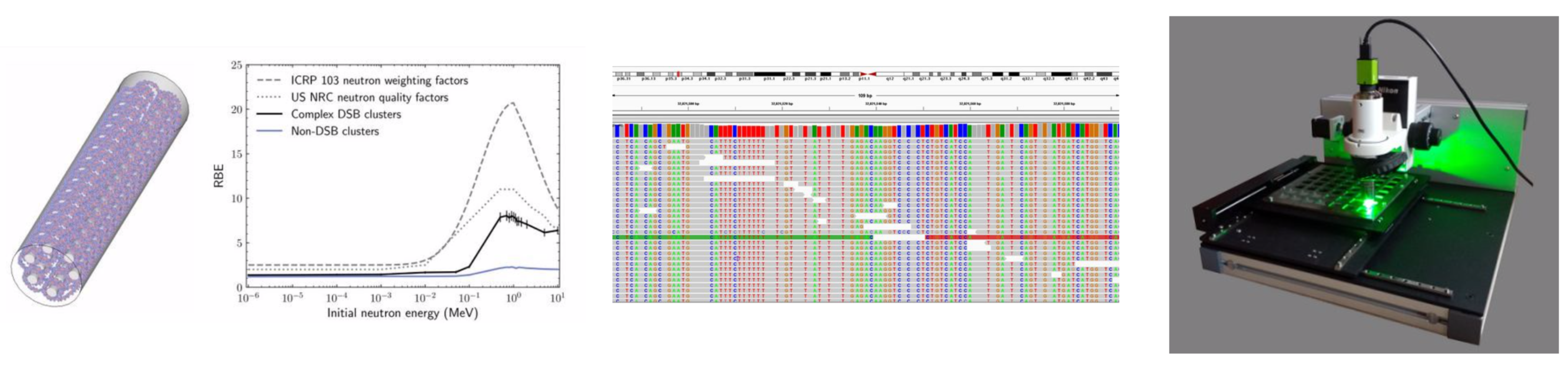 |
|---|
| Images related to NICE research projects (left to right): Monte Carlo DNA modelling, the ICRP radiation weighting factors for neutrons, single-cell DNA sequencing, track-structure neutron dosimetry using the TASL etch detector system. |
Rationale for program
Ionizing radiation is a carcinogen that is encountered in our natural and artificial environments. We cannot avoid it. Thankfully, radiological protection laws and regulations reduce the carcinogenic risk to individuals that excess ionizing radiation poses. However, while these measures guard against unjustified use of radiation, they do not apply to three distinct populations:
- Radiotherapy and radiology patients, for whom the out-of-field radiation dose is limited only by the justification (i.e. prescription) of the medical practitioner,
- Astronauts travelling in deep space who will be exposed to a chronic smorgasbord of cosmic rays, and
- Individuals who may be exposed to unknown radiation levels in the unlikely, but not impossible, event of a nuclear accident, atomic bomb, or terrorist attack involving an improvised nuclear device.
Therefore, any improvement to our understanding of radiation-induced carcinogenesis will help clinicians, policy-makers, and first responders make informed decisions to better protect these populations. The NICE research program is exploiting the energy-dependence of the neutron radiation weighting factors for stochastic effects to help explain the biophysics of radiation-induced carcinogenesis.
Available NICE Project 1: DNA Repair Modelling
It is believed that ionizing radiation damages DNA through direct and indirect action. In direct action, the DNA itself is ionized, whereas in indirect action, the chemical species produced by the radiolysis of water in the vicinity of the DNA produce the damage. But evolutionary forces have ensured that DNA is very capable of repairing itself such that not all DNA damage is mutagenic. Therefore, in this project we will expand on previous work by the Kildea lab and incorporate DNA repair modelling into existing Monte Carlo models of direct and indirect radiation action.
Specifically, this project will examine the relative biological effectiveness (RBE) of neutrons to produce mutations in DNA, accounting for direct and indirect damage and DNA repair. Findings from the project will be compared with mutations measured experimentally using single-cell whole genome DNA sequencing of irradiated cells.
This project will build upon the PhD thesis and the published journal article of Logan Montgomery that modelled direct action DNA damage, the MSc thesis of James Manalad that modelled indirect action DNA damage (paper in preparation), and the MSc thesis and published journal article of Chris Lund. Previous projects were carried out using Geant4 and the TOPAS and TOPAS-nBio Monte Carlo modelling frameworks and the presently-proposed project will do so also. Modelling will be consistent with the experimental work of PhD student Felix Mathew who is irradiating cells and performing single-cell whole genome sequencing.
Available NICE Project 2: Etch detector usage and modelling
The spectrum of neutrons produced by a radiotherapy linac can be measured in air using a neutron spectrometer, and the Kildea lab has pioneered the use of a device known as the nested neutron spectrometer (NNS) in radiotherapy (see papers by Robert Maglieri, Logan Montgomery and Felix Mathew). But, the neutron spectrum experienced by a sample of irradiated cells is very different to the in-air neutron spectrum as neutrons quickly loose their energies when they encounter matter. Therefore, in order to properly understand how neutrons damage DNA, the neutron spectrum actually experienced by the cells must be determined. This can be done in two ways: (1) using Monte Carlo modelling, and (2) using in-vivo neutron spectrometry. In this project, we will do both. We will use the Geant4 MC package to model the irradiation of human lymphoblastoid cells by neutrons at a linac and we will use neutron track-etch detectors (CR-39 film) to determine the neutron spectrum experienced by the cells.
Radiation Oncology Knowledge Sharing
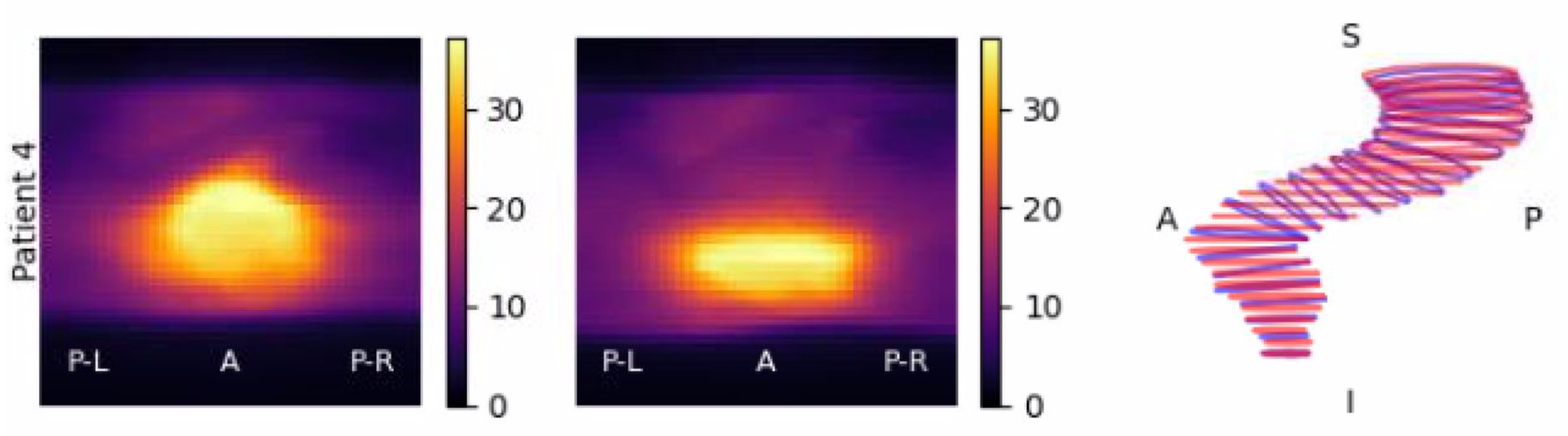 |
|---|
| The rtdsm dose-surface mapping software of Haley Patrick |
Rationale for the program
The ROKS research program is focused on improving the experiences and outcomes of radiotherapy patients by sharing knowledge through data.
Ongoing projects include:
- Generation of dose-surface maps to accumulate dose to the rectum during prostate radiotherapy,
- Prediction of pain in patients with bone metastases using radiomics and natural language processing, and
- Predicting when the radiotherapy of head and neck cancer patients should be replanned in order to account for anatomical changes that are anticipated over the course of treatment.
Opal and the Quebec SmartCare Consortium
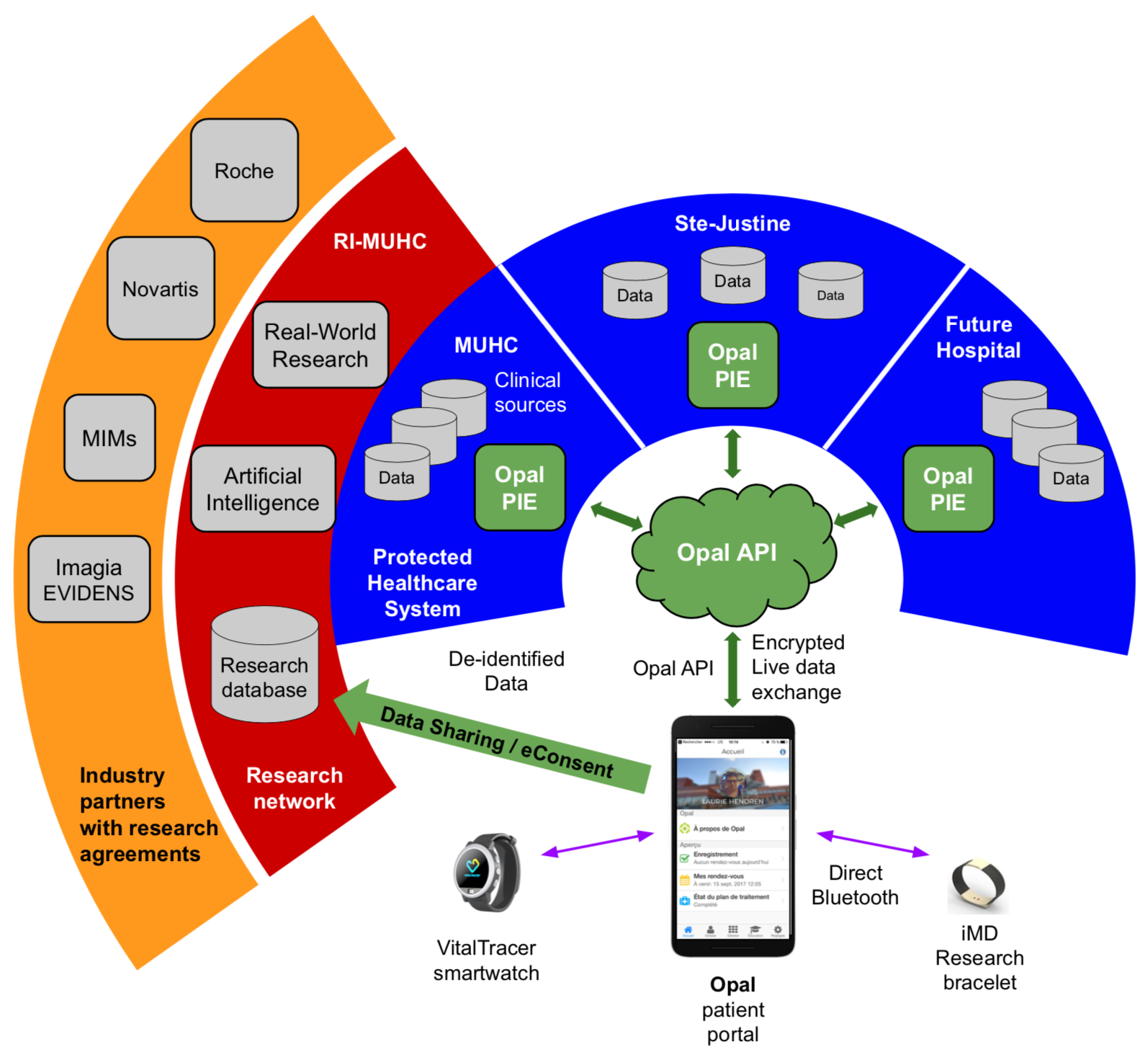 |
|---|
| The Quebec SmartCare Consortium is using the Opal patient portal as secure patient-controlled data-sharing hub to power real-world evidence research and a learning healthcare system in Quebec |
Rationale for the program
This research program aims to do two things:
- Empower patients with access to their healthcare data, and
- Inform real-world evidence research and develop a learning healthcare system through patient-controlled data sharing.
Randomized controlled clinical trials are considered the gold-standard for clinical knowledge generation and they form the bulk of clinical research. But, clinical trials are difficult to undertake, less than 5% of patients are enrolled in them, and due to their strict eligibility and rigid experimental design, many clinical trials are not representative of the real world. Therefore, a new form of clinical research, known as real-world evidence (RWE) research is gaining popularity. This form of research is retrospective in nature and generates evidence regarding clinical effectiveness using artificial intelligence techniques applied to real-world data from electronic medical records and from patients themselves.
The Quebec SmartCare Consortium (QSCC) is a group of public and private partners in Quebec that have come together to use the data-sharing capabilities of the Opal patient portal to power real-world evidence research and a learning healthcare system in Quebec.
Available Opal and QSCC Project 1: New and standardized data in oncology
This project forms part of the QSCC.
A lot of cancer patient data (demographics, disease information, treatment information) are available in our radiation oncology information system (Aria by Varian Medical Systems). And, through research projects approved by the McGill University Health Centre research ethics board, we have access to these data for research. But, many important data are not collected and many data types are not standardized. In particular, little or no data are collected regarding treatment outcomes and radiotherapy dose surface maps are not generated for hollow organs. Compounding this, a lot of the data that are collected are not standardized and so are difficult to aggregate. Therefore, this project will focus on:
- Generating evidence using data collected directly from patients themselves,
- The use of radiotherapy dose surface maps for hollow organs at risk, and
- Using the mCode data standard.
Patient-generated data will include:
- Electronic patient-reported outcomes and vital signs collected using the Opal patient portal
Vital signs will include:
- temperature,
- blood pressure,
- blood oxygen saturation,
- heart rate
- Weight
These data will be collected from patients in real-time during and after cancer treatment via smartwatches, smart bracelets, and smart weighing scales provided to patients and connected to the Opal patient patient portal app.
Dose surface maps will be generated using an open-source algorithm previously developed by Haley Patrick as part of her PhD project.
The mCode data standard will be incorporated into all work and will build on previous work by Kayla O’Sullivan and ongoing work by the CodeX FHIR accelerator at HL7 International.
This project will be a proof-of-principle feasibility initiative to show that it is possible to efficiently and accurately collect these new data and adhere to the mCode data standard.
Available Opal and QSCC Project 2: OncoBuddy
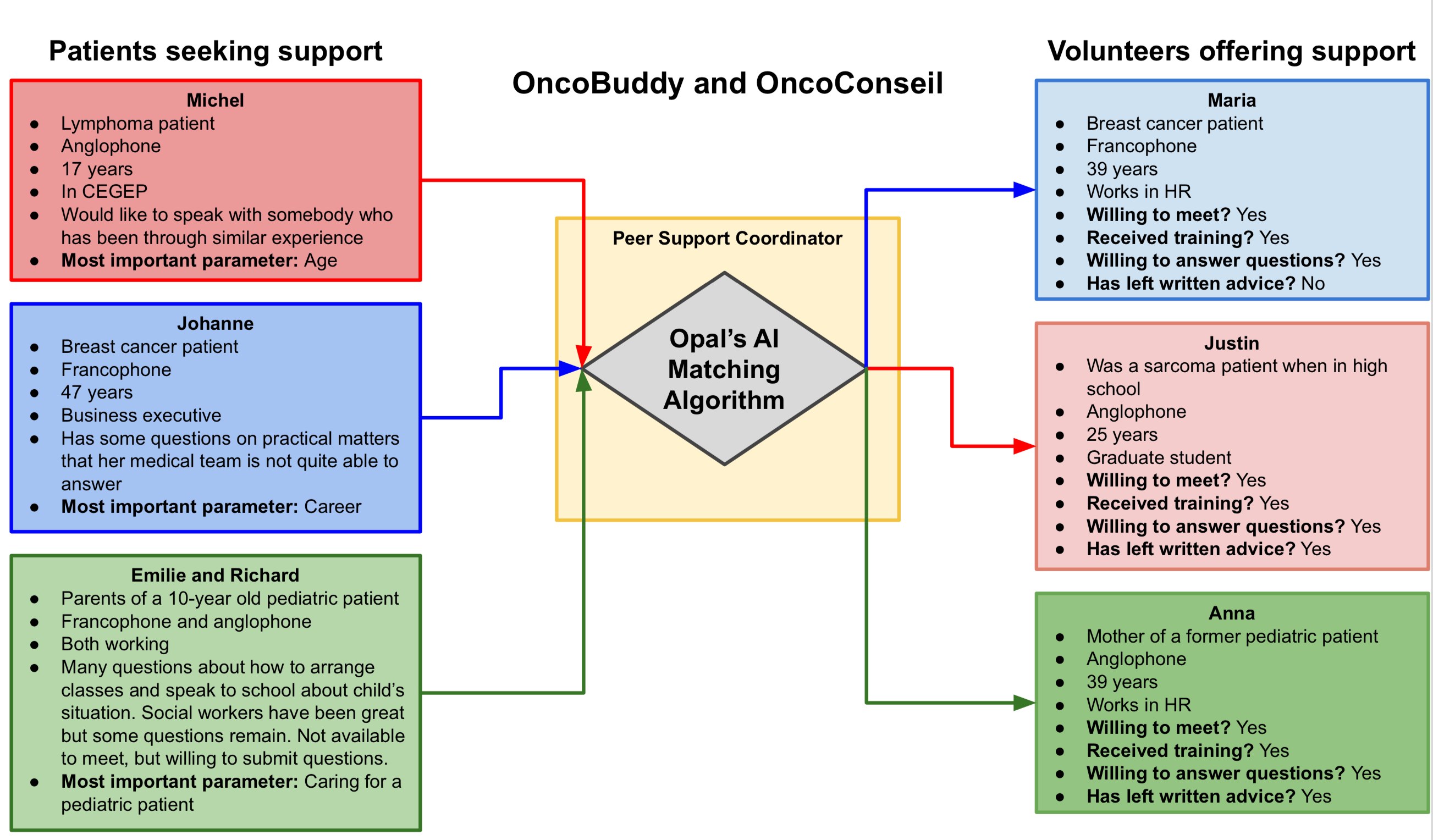 |
|---|
| The OncoBuddy and OncoConseil project will use AI techniques to match patients to each other for the purpose of peer support. |
The cancer experience is anxiety-provoking. As well as worrying about their disease, cancer patients (and their informal caregivers) must navigate many medical, logistical, and psychosocial challenges. The uncertainty of which challenge will come next, and how, can be distressing. But patients can help each other. One way in which cancer-related distress can be reduced is through peer support, in which a new cancer patient obtains some foresight through the hindsight of a previous patient. Therefore, in this research project, we will develop and pilot-test a novel, patient-suggested, artificial intelligence (AI) approach to improve the availability and the effectiveness of peer support in Quebec using the Opal patient portal app. In doing so we will help meet the informational and self-management needs of current patients by harnessing the expertise of previous patients.
While this project has a strong psychosocial motivation, it is grounded in quantitative oncology data and AI principles that are well suited to medical physics. Given the data and preferences of a new patient, what is the most appropriate way to identify those previous patients that best match the preferences of the new patient? And, how can the matching algorithm continuously learn and improve itself?
The new dimension of psychosocial data to be collected from patients in this initiative will ultimately enrich the body of real-world data for real-world evidence research. For example, currently no data are collected in healthcare regarding patient lifestyles, religious preferences, and cultural preferences, all of which may ultimately impact cancer treatment outcomes.
This project will build on previous work undertaken by Aixa Andrade who investigated various matching algorithms (stable marriage, hospital residence, stable roommate, genetic algorithms) and recommendation systems (nearest neighbors, relevance feedback, etc). It will be guided by a stakeholder co-design committee of patients, clinicians, and researchers that the student will become part of, and it will be coupled with the digital twinning PhD project of Kayla O’Sullivan. The student will also have the opportunity to work on the Opal patient portal software.
Ongoing Student Projects
Below is a list of ongoing student projects in the Kildea lab related to the NICE, ROKS and Opal research programs.