Monte Carlo modeling to investigate the suitability of using single-cell DNA sequencing of irradiated cells in the mutation signature analysis of low- and high-LET radiation
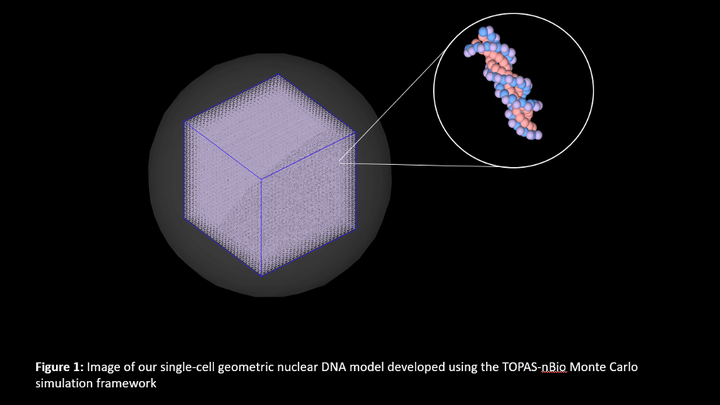 Image credit: Felix Mathew
Image credit: Felix MathewAbstract
Purpose: Behjati et al. (2016) has shown that radiotherapy-associated second cancers exhibit mutation signatures (MS) specific to ionizing radiation. Based on the Behjati results, we expect distinct MS for low- and high-LET radiations but stratifying radiotherapy-associated cancers by LET is an arduous task. We hypothesize that cells irradiated in vitro can serve as an alternative to tumor cells to identify radiation-induced MS by performing DNA sequencing at the single-cell level. With this in mind, we used Monte Carlo modeling to investigate if low- and high-LET radiation introduces distinguishable DNA damage patterns in cells’ genomes. Methods: Our single-cell geometric nuclear DNA model, developed using the TOPAS-nBio toolkit, was exposed to low-LET photons and high-LET neutrons of various doses. Direct and indirect radiation actions were tracked for all secondary particles, including reactive oxygen species. Various types of DNA damage were scored with repeated simulations. Results: We observed a significant increase in the number of radiation-induced DNA damage sites with dose for both radiation qualities that should be experimentally measurable using single-cell DNA sequencing. On comparing, we found that the number of base lesions was significantly higher for low-LET photons than for neutrons, whereas the number of clustered DNA damages was higher for neutrons than photons. Conclusion: Our simulations show that the DNA damage induced by low- and high-LET radiations should be distinguishable in a cell population using single-cell sequencing.